Structural Similarity Index Measure (SSIM)¶
Module Interface¶
- class torchmetrics.image.StructuralSimilarityIndexMeasure(gaussian_kernel=True, sigma=1.5, kernel_size=11, reduction='elementwise_mean', data_range=None, k1=0.01, k2=0.03, return_full_image=False, return_contrast_sensitivity=False, **kwargs)[source]¶
Compute Structural Similarity Index Measure (SSIM).
As input to
forwardandupdatethe metric accepts the following inputAs output of forward and compute the metric returns the following output
ssim(Tensor): ifreduction!='none'returns float scalar tensor with average SSIM value over sample else returns tensor of shape(N,)with SSIM values per sample
- Parameters:
preds¶ – estimated image
target¶ – ground truth image
gaussian_kernel¶ (
bool) – IfTrue(default), a gaussian kernel is used, ifFalsea uniform kernel is usedsigma¶ (
Union[float,Sequence[float]]) – Standard deviation of the gaussian kernel, anisotropic kernels are possible. Ignored if a uniform kernel is usedkernel_size¶ (
Union[int,Sequence[int]]) – the size of the uniform kernel, anisotropic kernels are possible. Ignored if a Gaussian kernel is usedreduction¶ (
Literal['elementwise_mean','sum','none',None]) –a method to reduce metric score over individual batch scores
'elementwise_mean': takes the mean'sum': takes the sum'none'orNone: no reduction will be applied
data_range¶ (
Union[float,tuple[float,float],None]) – the range of the data. If None, it is determined from the data (max - min). If a tuple is provided then the range is calculated as the difference and input is clamped between the values.return_full_image¶ (
bool) – If true, the fullssimimage is returned as a second argument. Mutually exclusive withreturn_contrast_sensitivityreturn_contrast_sensitivity¶ (
bool) – If true, the constant term is returned as a second argument. The luminance term can be obtained with luminance=ssim/contrast Mutually exclusive withreturn_full_imagekwargs¶ (
Any) – Additional keyword arguments, see Advanced metric settings for more info.
Example
>>> import torch >>> from torchmetrics.image import StructuralSimilarityIndexMeasure >>> preds = torch.rand([3, 3, 256, 256]) >>> target = preds * 0.75 >>> ssim = StructuralSimilarityIndexMeasure(data_range=1.0) >>> ssim(preds, target) tensor(0.9219)
- plot(val=None, ax=None)[source]¶
Plot a single or multiple values from the metric.
- Parameters:
val¶ (
Union[Tensor,Sequence[Tensor],None]) – Either a single result from calling metric.forward or metric.compute or a list of these results. If no value is provided, will automatically call metric.compute and plot that result.ax¶ (
Optional[Axes]) – An matplotlib axis object. If provided will add plot to that axis
- Return type:
- Returns:
Figure and Axes object
- Raises:
ModuleNotFoundError – If matplotlib is not installed
>>> # Example plotting a single value >>> import torch >>> from torchmetrics.image import StructuralSimilarityIndexMeasure >>> preds = torch.rand([3, 3, 256, 256]) >>> target = preds * 0.75 >>> metric = StructuralSimilarityIndexMeasure(data_range=1.0) >>> metric.update(preds, target) >>> fig_, ax_ = metric.plot()
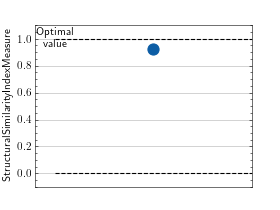
>>> # Example plotting multiple values >>> import torch >>> from torchmetrics.image import StructuralSimilarityIndexMeasure >>> preds = torch.rand([3, 3, 256, 256]) >>> target = preds * 0.75 >>> metric = StructuralSimilarityIndexMeasure(data_range=1.0) >>> values = [ ] >>> for _ in range(10): ... values.append(metric(preds, target)) >>> fig_, ax_ = metric.plot(values)
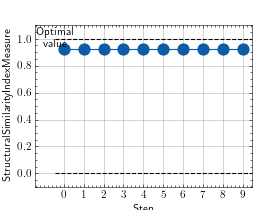
Functional Interface¶
- torchmetrics.functional.image.structural_similarity_index_measure(preds, target, gaussian_kernel=True, sigma=1.5, kernel_size=11, reduction='elementwise_mean', data_range=None, k1=0.01, k2=0.03, return_full_image=False, return_contrast_sensitivity=False)[source]¶
Compute Structural Similarity Index Measure.
- Parameters:
gaussian_kernel¶ (
bool) – If true (default), a gaussian kernel is used, if false a uniform kernel is usedsigma¶ (
Union[float,Sequence[float]]) – Standard deviation of the gaussian kernel, anisotropic kernels are possible. Ignored if a uniform kernel is usedkernel_size¶ (
Union[int,Sequence[int]]) – the size of the uniform kernel, anisotropic kernels are possible. Ignored if a Gaussian kernel is usedreduction¶ (
Literal['elementwise_mean','sum','none',None]) –a method to reduce metric score over labels.
'elementwise_mean': takes the mean'sum': takes the sum'none'orNone: no reduction will be applied
data_range¶ (
Union[float,tuple[float,float],None]) – the range of the data. If None, it is determined from the data (max - min). If a tuple is provided then the range is calculated as the difference and input is clamped between the values.return_full_image¶ (
bool) – If true, the fullssimimage is returned as a second argument. Mutually exclusive withreturn_contrast_sensitivityreturn_contrast_sensitivity¶ (
bool) – If true, the constant term is returned as a second argument. The luminance term can be obtained with luminance=ssim/contrast Mutually exclusive withreturn_full_image
- Return type:
- Returns:
Tensor with SSIM score
- Raises:
TypeError – If
predsandtargetdon’t have the same data type.ValueError – If
predsandtargetdon’t haveBxCxHxW shape.ValueError – If the length of
kernel_sizeorsigmais not2.ValueError – If one of the elements of
kernel_sizeis not anodd positive number.ValueError – If one of the elements of
sigmais not apositive number.
Example
>>> from torchmetrics.functional.image import structural_similarity_index_measure >>> preds = torch.rand([3, 3, 256, 256]) >>> target = preds * 0.75 >>> structural_similarity_index_measure(preds, target) tensor(0.9219)
